Single Cell Analysis Facility
Overview
The Chromium Single Cell Solution delivers a scalable microfluidic platform for digital gene expression profiling of up to 240,000 cells per run (8 wells). The technology permits high throughput single cell transcriptomics from a variety of cell types (up to 50 μm in diameter) as well as single-nuclei profiling using 10X Genomics approved protocols. The 10x GemCode Technology samples a pool of ~750,000 barcodes to separately index the transcriptome of each cell. It does so by partitioning thousands of cells into nanoliter-scale Gel Bead-In-Emulsions (GEMs) where all generated cDNAs share a common barcode. Libraries are generated and sequenced from the cDNA and the barcodes are then used to associate individual reads back to the individual partitions. Up to 8 samples can be processed in parallel in the Chromium and multiplexing option adds many more cells to the mix. More information about Chromium technology and technical resources can be found on the 10X Genomics Website.
NEW
We now have received Chromium iX which enables single cell sequencing from PFA-Fixed and FFPE in addition to fresh and fresh frozen cells and Nuclei.
Contact us to discuss your project!
With the RNA-Flex protocol up to 16 samples can be pooled in a single well and over 1 million cells can be targeted on a single Chromium iX run.
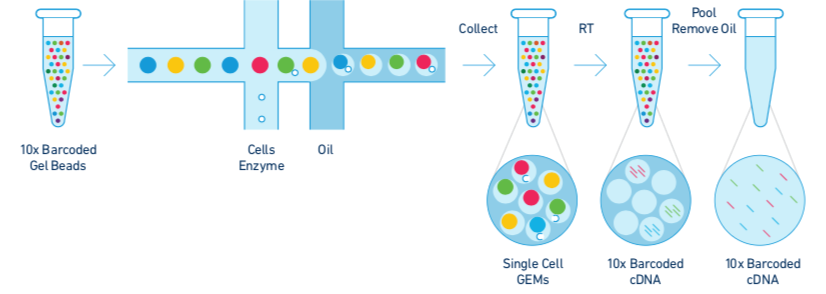
Chromium Single Cell Solution. Formation of GEMs. RT reaction takes place inside each GEM, which is then pooled for cDNA amplification and library construction in bulk.
Applications
- Single Cell RNA Flex (Fixed RNA Profiling)
- Single cell 3’ RNA-seq
- Single cell 3’ RNA-seq for Cell Mutiplexing
- Single Cell Multiome ATAC+ GEX
- Single Cell 3' RNA-Seq for CRISPR Screening
- Visium Spatial Gene Expression
- Single cell immune profiling
- Single Cell ATAC-seq
- Single Cell CNVs
Services
- Cell counts and viability assessment on the Countess II
- Cell washing and resuspension as required for input into the Chromium Single Cell Controller.
- Prob Hybridization for RNA Flex Assay
- Cell capture and GEM formation
- cDNA amplification
- Library preparation with quality assurance
- Next generation sequencing on the Illumina MiniSeq, NextSeq 500 or NextSeq 2000 as required to achieve desired depth
- Delivery of FastQ files and QC results
- Data files can be analyzed with 10X Genomics free software solutions Cell Ranger and Cell Loupe Browser.
- Optional data analysis services can also be provided by the OHRI Bioinformatics Facility
Consultation
- Experimental Design
- Pilot Projects
- Proof of Concept Experiments
- Grant Support
- Manuscript Preparation
- Full support necessary to bring your project to successful completion
Instrumentation
Countess II Cell Counter and Viability Analyzer

Luna FL, Dual Fluorescence Cell Counter
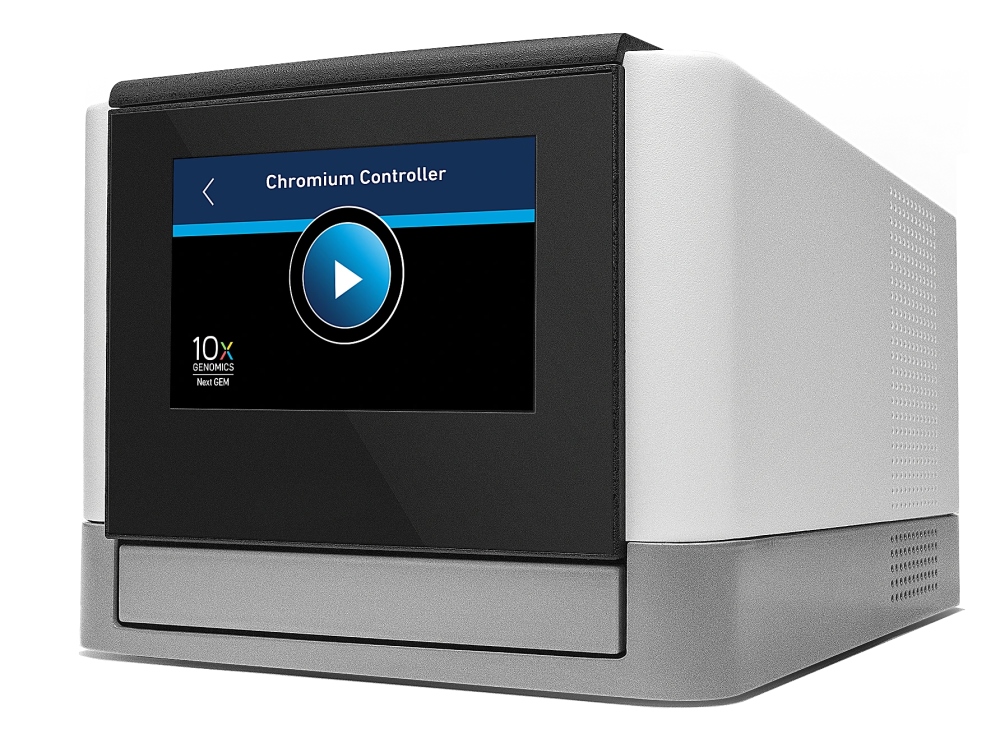
10X Genomics Chromium Single Cell Controller

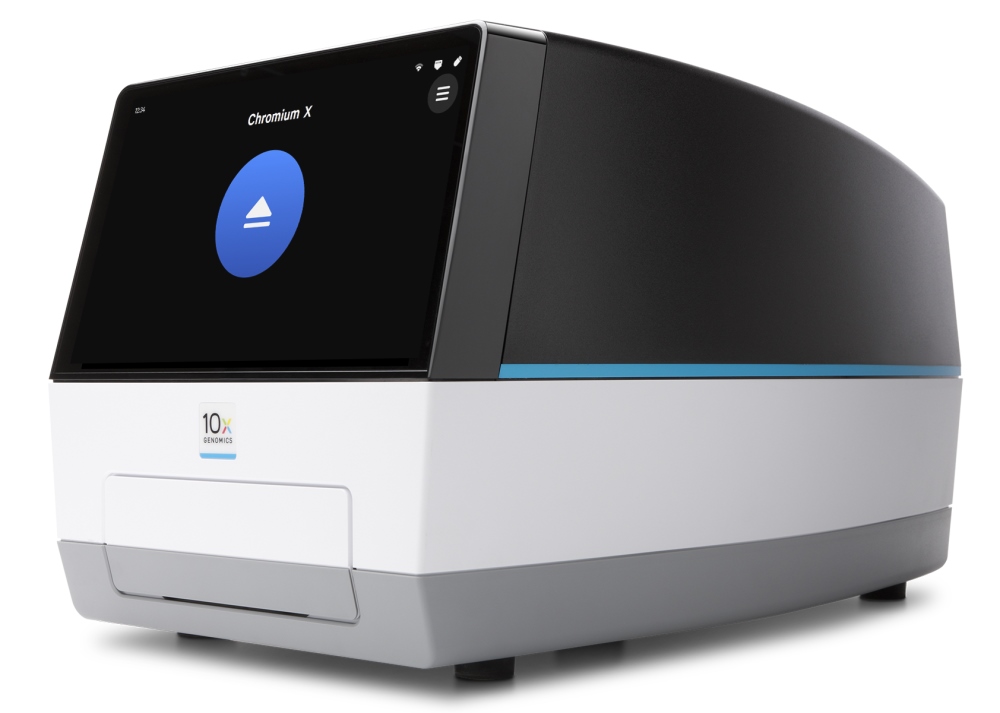
10x Genomics iX Chromium Single Cell Controller

General Recommendations
- To initiate a project, please contact Kathy
Sheikheleslamy or Damian Carragher
- Please refer to the Single Cell Protocols Cell Preparation Guide for recommended
best practices for washing, counting, and concentrating cells.
- Please refer to the Fixation of Cells and Nuclei for Chromium Fixed RNA Profiling Protocol for recommended steps of sample prep.
- A service agreement must be established and signed by the PI and Institutional Signing Authority (external clients) for each project prior to processing samples. The agreement will include a statement of work and a quotation.
- All Single Cell service requests must be entered in iLab/ Infinity Idea Elan and reference the Service Agreement number provided when the project is initiated. Samples should be provided with a copy of the sample submission form and a reference to the request ID
- Each sample should be submitted as a single cell suspension at a concentration of 1,000-1,500 cells/ul in compatible buffer/ media at >90% viability.
- Clients will provide approval of cell counting and viability assessment on the Luna/ Countess II to proceed with the experiment.
- Cells should be received no later than 1:00 pm on previously scheduled analysis date.
Pricing
The cost of this service is dependent on several variables including the assay, number of samples analyzed at the same time, number of cells targeted per sample, and the read depth per cell.
Please contact us for a consultation so that we can determine the best assay and approach for your specific project and budget.
Contact Information
Kathy Sheikheleslamy
ksheikheleslamy@ohri.ca
613.737.8899 x73251
Damian Carragher
dcarragher@ohri.ca
613.737.8899 x73110
